Profile data processing
Goal: You want to find all peaks in your profile data.
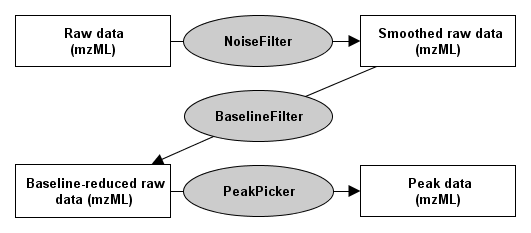
The first step shown here is the elimination of noise using a NoiseFilter. The now smoothed profile data can be further processed by subtracting the baseline with the BaselineFilter. Then use one of the PeakPickers to find all peaks in the baseline-reduced profile data.
We offer two different smoothing filters: NoiseFilterGaussian and NoiseFilterSGolay. If you want to use the Savitzky Golay filter, or our BaselineFilter with non equally spaced profile data, e.g. TOF data, you have to generate equally spaced data using the Resampler tool.
Picking peaks with a PeakPicker
The PeakPicker tools allow for picking peaks in profile data. Currently, there are two different TOPP tools available, PeakPickerWavelet and PeakPickerHiRes.
| PeakPickerWavelet Input data: profile data (low/medium resolution) |
Description:
This peak picking algorithm uses the continuous wavelet transform of a raw data signal to detect mass peaks. Afterwards a given asymmetric peak function is fitted to the raw data and important peak parameters (e.g. fwhm) are extracted. In an optional step these parameters can be optimized using a non-linear optimization method.
The algorithm is described in detail in Lange et al. (2006) Proc. PSB-06.
Application:
This algorithm was designed for low and medium resolution data. It can also be applied to high-resolution data, but can be slow on large datasets.
See the PeakPickerCWT class documentation for a parameter list.
|
| PeakPickerHiRes Input data: profile data (high resolution) |
Description:
This peak-picking algorithm detects ion signals in raw data and reconstructs the corresponding peak shape by cubic spline interpolation. Signal detection depends on the signal-to-noise ratio which is adjustable by the user (see parameter signal_to_noise). A picked peak's m/z and intensity value is given by the maximum of the underlying peak spline. Please notice that this method is still experimental since it has not been tested thoroughly yet.
Application:
The algorithm is best suited for high-resolution MS data (FT-ICR-MS, Orbitrap). In high-resolution data, the signals of ions with similar mass-to-charge ratios (m/z) exhibit little or no overlapping and therefore allow for a clear separation. Furthermore, ion signals tend to show well-defined peak shapes with narrow peak width. These properties facilitate a fast computation of picked peaks so that even large data sets can be processed very quickly.
See the PeakPickerHiRes class documentation for a parameter list.
|
Finding the right parameters for the
NoiseFilters, the BaselineFilter and the PeakPickers
Finding the right parameters is not trivial. The default parameters will not work on most datasets. In order to find good parameters, we propose the following procedure:
- Load the data in TOPPView
- Extract a single scan from the middle of the HPLC gradient (Right click on scan)
- Experiment with the parameters until you have found the proper settings
- You can find the NoiseFilters, the BaselineFilter, and the PeakPickers in TOPPView in the menu 'Layer' - 'Apply TOPP tool'


 1.8.16
1.8.16